Browse by Organisms Motifs
| M. musculus Motifs of MC11 group | ||||
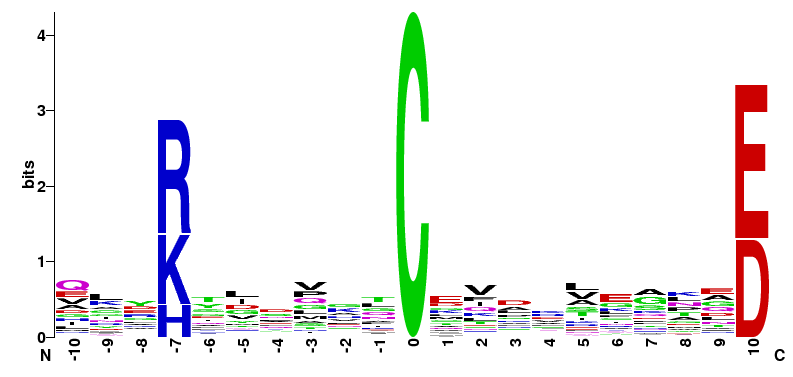 |
| No. | SwissProt ID | Position | S-nitrosylated Peptide | PubMed ID |
| 1 | 1433T_MOUSE | 25 | QAERYDDMAT C MKAVTEQGAE | 24926564 |
| 2 | 1433T_MOUSE | 25 | QAERYDDMAT C MKAVTEQGAE | 20925432 |
| 3 | 1433Z_MOUSE | 25 | QAERYDDMAA C MKSVTEQGAE | 24926564 |
| 4 | 1433Z_MOUSE | 25 | QAERYDDMAA C MKSVTEQGAE | 20925432 |
| 5 | 1433Z_MOUSE | 25 | QAERYDDMAA C MKSVTEQGAE | 21278135 |
| 6 | 1433Z_MOUSE | 25 | QAERYDDMAA C MKSVTEQGAE | 22865876 |
| 7 | AATM_MOUSE | 187 | QGYRYYDPKT C GFDFSGALED | 22178444 |
| 8 | AATM_MOUSE | 187 | QGYRYYDPKT C GFDFSGALED | 22588120 |
| 9 | AATM_MOUSE | 187 | QGYRYYDPKT C GFDFSGALED | 24895380 |
| 10 | AATM_MOUSE | 187 | QGYRYYDPKT C GFDFSGALED | 19483679 |
| 11 | AATM_MOUSE | 187 | QGYRYYDPKT C GFDFSGALED | 20925432 |
| 12 | AATM_MOUSE | 187 | QGYRYYDPKT C GFDFSGALED | 21278135 |
| 13 | AATM_MOUSE | 187 | QGYRYYDPKT C GFDFSGALED | 20837516 |
| 14 | AATM_MOUSE | 187 | QGYRYYDPKT C GFDFSGALED | 22865876 |
| 15 | ACAD8_MOUSE | 349 | QEEREDAVAL C SMAKLFATEE | 21278135 |
| 16 | ACAD8_MOUSE | 349 | QEEREDAVAL C SMAKLFATEE | 22865876 |
| 17 | ACSL5_MOUSE | 471 | TAGHVGTPVA C NFVKLEDVAD | 20837516 |
| 18 | ACSM5_MOUSE | 342 | TRYKFQCLRH C LTGGEALNPD | 22178444 |
| 19 | ACSM5_MOUSE | 342 | TRYKFQCLRH C LTGGEALNPD | 20837516 |
| 20 | AEDO_MOUSE | 225 | IRPKEASGSA C DLPREVWLLE | 21278135 |
| 21 | AL1A1_MOUSE | 186 | FIWKIGPALS C GNTVVVKPAE | 20837516 |
| 22 | AN32B_MOUSE | 123 | DCLKSLDLFG C EVTNRSDYRE | 20925432 |
| 23 | AT1A1_MOUSE | 211 | ADLRIISANG C KVDNSSLTGE | 24895380 |
| 24 | AT1A1_MOUSE | 211 | ADLRIISANG C KVDNSSLTGE | 21278135 |
| 25 | AT1A2_MOUSE | 209 | ADLRIISSHG C KVDNSSLTGE | 24895380 |
| 26 | AT1A2_MOUSE | 209 | ADLRIISSHG C KVDNSSLTGE | 21278135 |
| 27 | AT1A3_MOUSE | 201 | ADLRIISAHG C KVDNSSLTGE | 24895380 |
| 28 | AT2A2_MOUSE | 635 | GDNKGTAVAI C RRIGIFGQDE | 21278135 |
| 29 | AT2A2_MOUSE | 635 | GDNKGTAVAI C RRIGIFGQDE | 22865876 |
| 30 | BDH_MOUSE | 115 | DRLRTIQLNV C NSEEVEKAVE | 21278135 |
| 31 | BDH_MOUSE | 115 | DRLRTIQLNV C NSEEVEKAVE | 20837516 |
| 32 | BDH_MOUSE | 115 | DRLRTIQLNV C NSEEVEKAVE | 22865876 |
| 33 | BPHL_MOUSE | 234 | ICRHLLPLVQ C PTLIVHGEKD | 21278135 |
| 34 | BPHL_MOUSE | 234 | ICRHLLPLVQ C PTLIVHGEKD | 22865876 |
| 35 | C1QBP_MOUSE | 182 | TDGKKTLVLD C HYPEDEIGHE | 22865876 |
| 36 | CISD1_MOUSE | 83 | RCWRSKKFPF C DGAHIKHNEE | 21278135 |
| 37 | CISD1_MOUSE | 83 | RCWRSKKFPF C DGAHIKHNEE | 22865876 |
| 38 | COF1_MOUSE | 80 | TFVKMLPDKD C RYALYDATYE | 24926564 |
| 39 | COF1_MOUSE | 80 | TFVKMLPDKD C RYALYDATYE | 20925432 |
| 40 | COF1_MOUSE | 80 | TFVKMLPDKD C RYALYDATYE | 21278135 |
| 41 | COR1C_MOUSE | 330 | MPKRGLDVNK C EIARFFKLHE | 20925432 |
| 42 | CPPED_MOUSE | 54 | GLMKAWSTGN C DAGGDEWGQE | 21278135 |
| 43 | CPT1B_MOUSE | 462 | WFDKSFTLIS C KNGLLGLNTE | 21278135 |
| 44 | CPT1B_MOUSE | 462 | WFDKSFTLIS C KNGLLGLNTE | 22865876 |
| 45 | CTL2_MOUSE | 401 | AVYKVVDDTA C PLLRKTCNPE | 24895380 |
| 46 | CX6B1_MOUSE | 65 | EWYRRVYKSL C PVSWVSAWDD | 22588120 |
| 47 | CX6B1_MOUSE | 65 | EWYRRVYKSL C PVSWVSAWDD | 21278135 |
| 48 | CX6B1_MOUSE | 65 | EWYRRVYKSL C PVSWVSAWDD | 22865876 |
| 49 | DCMC_MOUSE | 205 | NLERVTWHSP C EVLQKISECE | 21278135 |
| 50 | DESM_MOUSE | 332 | EYRHQIQSYT C EIDALKGTND | 21278135 |
| 51 | DESM_MOUSE | 332 | EYRHQIQSYT C EIDALKGTND | 22865876 |
| 52 | EFTS_MOUSE | 239 | VLGKYGALVI C ETPEQIANLE | 21278135 |
| 53 | ES1_MOUSE | 219 | EAIKALGAKH C VKGVTEAHVD | 21278135 |
| 54 | ETHE1_MOUSE | 170 | GCGRTDFQQG C AKTLYHSVHE | 21278135 |
| 55 | GFAP_MOUSE | 291 | DYRRQLQALT C DLESLRGTNE | 22178444 |
| 56 | GFAP_MOUSE | 291 | DYRRQLQALT C DLESLRGTNE | 22588120 |
| 57 | GFAP_MOUSE | 291 | DYRRQLQALT C DLESLRGTNE | 24895380 |
| 58 | GFAP_MOUSE | 291 | DYRRQLQALT C DLESLRGTNE | 19101475 |
| 59 | GLRX3_MOUSE | 148 | RLKKLTHAAP C MLFMKGTPQE | 21278135 |
| 60 | GNAO_MOUSE | 108 | KERKTDSKMV C DVVSRMEDTE | 24895380 |
| 61 | HOP_MOUSE | 34 | VNKHPDPTTL C LIAAEAGLTE | 21278135 |
| 62 | HXK2_MOUSE | 517 | APVKMLPTYV C ATPDGTEKGD | 21278135 |
| 63 | ICAL_MOUSE | 408 | AKAKEERQEK C GEDEDTVPAE | 21278135 |
| 64 | IPO7_MOUSE | 757 | LQCKGRGIDQ C IPLFVEAALE | 21278135 |
| 65 | K0564_MOUSE | 857 | EADKAPTNVT C ILKTLVENGE | 21278135 |
| 66 | KPYM_MOUSE | 474 | HLYRGIFPVL C KDAVLNAWAE | 24895380 |
| 67 | KPYM_MOUSE | 474 | HLYRGIFPVL C KDAVLNAWAE | 20925432 |
| 68 | KPYM_MOUSE | 474 | HLYRGIFPVL C KDAVLNAWAE | 21278135 |
| 69 | KPYM_MOUSE | 474 | HLYRGIFPVL C KDAVLNAWAE | 22865876 |
| 70 | MP2K4_MOUSE | 244 | LLDRSGNIKL C DFGISGQLVD | 16418269 |
| 71 | MPPA_MOUSE | 465 | THSRKLPHEL C TLIRNVKPED | 21278135 |
| 72 | MT3_MOUSE | 38 | TNCKKSCCSC C PAGCEKCAKD | 22588120 |
| 73 | MT3_MOUSE | 38 | TNCKKSCCSC C PAGCEKCAKD | 24895380 |
| 74 | NDUBA_MOUSE | 77 | QYRRVPDITE C KEGDVLCIYE | 22865876 |
| 75 | NDUS1_MOUSE | 564 | CITRQDLPKD C FIVYQGHHGD | 21278135 |
| 76 | ODO1_MOUSE | 331 | VIRKELEQIF C QFDSKLEAAD | 21278135 |
| 77 | ODPB_MOUSE | 249 | VVAHSRPVGH C LEAAAVLSKE | 22865876 |
| 78 | OPA1_MOUSE | 786 | MYWKNRTQEQ C VHNETKNELE | 21278135 |
| 79 | OPA1_MOUSE | 786 | MYWKNRTQEQ C VHNETKNELE | 22865876 |
| 80 | PARK7_MOUSE | 106 | ESRKGLIAAI C AGPTALLAHE | 22865876 |
| 81 | PECR_MOUSE | 43 | AVSRELLHLG C NVVIASRKLD | 22178444 |
| 82 | PECR_MOUSE | 43 | AVSRELLHLG C NVVIASRKLD | 20837516 |
| 83 | PEX19_MOUSE | 229 | QQQHSVMVKI C EQFEAETPTD | 21278135 |
| 84 | PNPH_MOUSE | 142 | IRDHINLPGF C GQNPLRGPND | 20925432 |
| 85 | PSME4_MOUSE | 1343 | GRDKFSPRRF C LFKGIFRNFD | 21278135 |
| 86 | RB3GP_MOUSE | 181 | HKWRRMYMGE C QGPGVRTDFE | 21278135 |
| 87 | RBX2_MOUSE | 61 | AICRVQVMDA C LRCQAENKQE | 21278135 |
| 88 | REEP5_MOUSE | 120 | YMLKCGFLLW C MAPSPANGAE | 22865876 |
| 89 | RM47_MOUSE | 89 | EKVKSGASWT C QQLRNKSNED | 21278135 |
| 90 | ROA1_MOUSE | 175 | QKYHTVNGHN C EVRKALSKQE | 20925432 |
| 91 | ROA1_MOUSE | 175 | QKYHTVNGHN C EVRKALSKQE | 21278135 |
| 92 | ROA3_MOUSE | 196 | QKYHTINGHN C EVKKALSKQE | 20925432 |
| 93 | ROA3_MOUSE | 196 | QKYHTINGHN C EVKKALSKQE | 21278135 |
| 94 | RPN1_MOUSE | 546 | SRLKTEGSDL C DRVSEMQKLD | 22178444 |
| 95 | RPN1_MOUSE | 546 | SRLKTEGSDL C DRVSEMQKLD | 20837516 |
| 96 | RS27A_MOUSE | 145 | HFDRHYCGKC C LTYCFNKPED | 24926564 |
| 97 | RS27A_MOUSE | 145 | HFDRHYCGKC C LTYCFNKPED | 20925432 |
| 98 | RS28_MOUSE | 27 | VLGRTGSQGQ C TQVRVEFMDD | 22178444 |
| 99 | RS28_MOUSE | 27 | VLGRTGSQGQ C TQVRVEFMDD | 19101475 |
| 100 | RS28_MOUSE | 27 | VLGRTGSQGQ C TQVRVEFMDD | 20925432 |
| 101 | RS28_MOUSE | 27 | VLGRTGSQGQ C TQVRVEFMDD | 21278135 |
| 102 | RS28_MOUSE | 27 | VLGRTGSQGQ C TQVRVEFMDD | 22865876 |
| 103 | SAC1_MOUSE | 84 | VITKKMKVGE C FNHAVWRATD | 20925432 |
| 104 | SAR1A_MOUSE | 178 | LNARPMEVFM C SVLKRQGYGE | 21278135 |
| 105 | SAR1B_MOUSE | 178 | LNARPLEVFM C SVLKRQGYGE | 21278135 |
| 106 | SPF45_MOUSE | 343 | ECEKYGKVGK C VIFEIPGAPD | 20925432 |
| 107 | SYEP_MOUSE | 856 | PKAKVTEAVE C LLSLKAEYKE | 20837516 |
| 108 | SYRC_MOUSE | 86 | INSRLQEVFG C AIRAAYPDLE | 21278135 |
| 109 | SYRC_MOUSE | 369 | DGRKIVFVPG C SVPLTIVKSD | 21278135 |
| 110 | VIME_MOUSE | 328 | EYRRQVQSLT C EVDALKGTNE | 20925432 |
| 111 | ZC12C_MOUSE | 126 | KTHRQLCRSP C LEPRLLKHSD | 22865876 |
